The BioNJ distance method. More...
#include <Bpp/Phyl/Distance/BioNJ.h>
 Inheritance diagram for bpp::BioNJ:
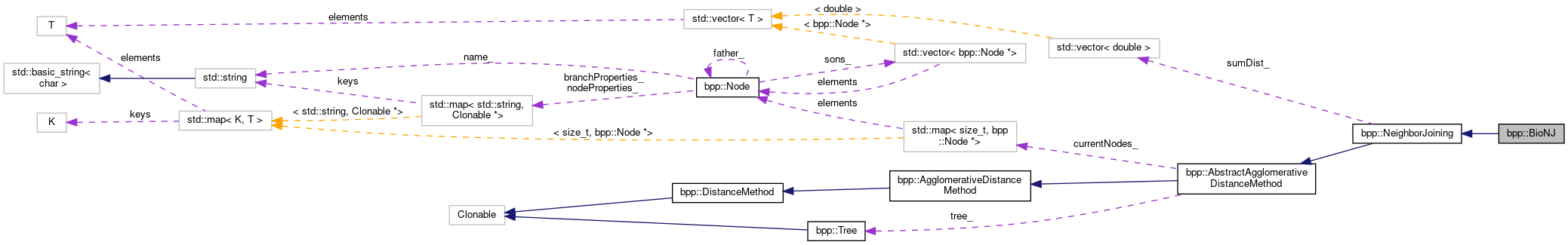
Inheritance diagram for bpp::BioNJ: Collaboration diagram for bpp::BioNJ:
Collaboration diagram for bpp::BioNJ:Public Member Functions | |
| BioNJ (bool rooted=false, bool positiveLengths=false, bool verbose=true) | |
| Create a new BioNJ object instance and compute a tree from a distance matrix. More... | |
| BioNJ (const DistanceMatrix &matrix, bool rooted=false, bool positiveLengths=false, bool verbose=true) throw (Exception) | |
| Create a new BioNJ object instance and compute a tree from a distance matrix. More... | |
| BioNJ * | clone () const |
| virtual | ~BioNJ () |
| std::string | getName () const |
| void | setDistanceMatrix (const DistanceMatrix &matrix) throw (Exception) |
| Set the distance matrix to use. More... | |
| void | computeTree () throw (Exception) |
| Compute the tree corresponding to the distance matrix. More... | |
| double | computeDistancesFromPair (const std::vector< size_t > &pair, const std::vector< double > &branchLengths, size_t pos) |
| Actualizes the distance matrix according to a given pair and the corresponding branch lengths. More... | |
| virtual void | outputPositiveLengths (bool yn) |
| virtual TreeTemplate< Node > * | getTree () const |
| Get the computed tree, if there is one. More... | |
| void | setVerbose (bool yn) |
| bool | isVerbose () const |
Protected Member Functions | |
| std::vector< size_t > | getBestPair () throw (Exception) |
| Get the best pair of nodes to agglomerate. More... | |
| std::vector< double > | computeBranchLengthsForPair (const std::vector< size_t > &pair) |
| Compute the branch lengths for two nodes to agglomerate. More... | |
| void | finalStep (int idRoot) |
| Method called when there ar eonly three remaining node to agglomerate, and creates the root node of the tree. More... | |
Specific methods. | |
| virtual Node * | getLeafNode (int id, const std::string &name) |
| Get a leaf node. More... | |
| virtual Node * | getParentNode (int id, Node *son1, Node *son2) |
| Get an inner node. More... | |
Protected Attributes | |
| std::vector< double > | sumDist_ |
| bool | positiveLengths_ |
| DistanceMatrix | matrix_ |
| Tree * | tree_ |
| std::map< size_t, Node * > | currentNodes_ |
| bool | verbose_ |
| bool | rootTree_ |
Private Attributes | |
| DistanceMatrix | variance_ |
| double | lambda_ |
Detailed Description
The BioNJ distance method.
Reference: Gascuel O. BIONJ: an improved version of the NJ algorithm based on a simple model of sequence data. Mol Biol Evol. 1997 Jul;14(7):685-95.
Constructor & Destructor Documentation
◆ BioNJ() [1/2]
|
inline |
Create a new BioNJ object instance and compute a tree from a distance matrix.
- Parameters
-
rooted Tell if the output tree should be rooted. positiveLengths Tell if negative lengths should be avoided. verbose Allow to display extra information, like progress bars.
Definition at line 70 of file BioNJ.h.
Referenced by clone().
◆ BioNJ() [2/2]
|
inline | ||||||||||||||||||||||||||||
Create a new BioNJ object instance and compute a tree from a distance matrix.
- Parameters
-
matrix Input distance matrix. rooted Tell if the output tree should be rooted. positiveLengths Tell if negative lengths should be avoided. verbose Allow to display extra information, like progress bars.
Definition at line 83 of file BioNJ.h.
References computeTree(), bpp::NeighborJoining::outputPositiveLengths(), and setDistanceMatrix().
◆ ~BioNJ()
Member Function Documentation
◆ clone()
|
inline |
◆ computeBranchLengthsForPair()
|
protectedvirtualinherited |
Compute the branch lengths for two nodes to agglomerate.
This method compute l1 and l2 given N1 and N2.
- Parameters
-
pair The indices of the nodes to be agglomerated.
- Returns
- A size 2 vector with branch lengths.
Implements bpp::AbstractAgglomerativeDistanceMethod.
Definition at line 92 of file NeighborJoining.cpp.
◆ computeDistancesFromPair()
|
virtual |
Actualizes the distance matrix according to a given pair and the corresponding branch lengths.
- Parameters
-
pair The indices of the nodes to be agglomerated. branchLengths The corresponding branch lengths. pos The index of the node whose distance ust be updated.
- Returns
- The distance between the 'pos' node and the agglomerated pair.
Implements bpp::AbstractAgglomerativeDistanceMethod.
◆ computeTree()
|
virtual | |||||||||||||
Compute the tree corresponding to the distance matrix.
This method implements the following algorithm: 1) Build all leaf nodes (getLeafNode method) 2) Get the best pair to agglomerate (getBestPair method) 3) Compute the branch lengths for this pair (computeBranchLengthsForPair method) 4) Build the parent node of the pair (getParentNode method) 5) For each remaining node, update distances from the pair (computeDistancesFromPair method) 6) Return to step 2 while there are more than 3 remaining nodes. 7) Perform the final step, and send a rooted or unrooted tree.
Reimplemented from bpp::AbstractAgglomerativeDistanceMethod.
Definition at line 60 of file BioNJ.cpp.
References bpp::Node::setDistanceToFather().
Referenced by BioNJ(), bpp::TreeTools::MRP(), and bpp::TreeTools::MRPMultilabel().
◆ finalStep()
|
protectedvirtualinherited |
Method called when there ar eonly three remaining node to agglomerate, and creates the root node of the tree.
- Parameters
-
idRoot The id of the root node.
Implements bpp::AbstractAgglomerativeDistanceMethod.
Definition at line 117 of file NeighborJoining.cpp.
References bpp::Node::addSon(), and bpp::Node::setDistanceToFather().
◆ getBestPair()
|
protectedvirtualinherited | |||||||||||||
Get the best pair of nodes to agglomerate.
Define the criterion to chose the next pair of nodes to agglomerate. This criterion uses the matrix_ distance matrix.
- Returns
- A size 2 vector with the indices of the nodes.
- Exceptions
-
Exception If an error occured.
Implements bpp::AbstractAgglomerativeDistanceMethod.
Definition at line 51 of file NeighborJoining.cpp.
◆ getLeafNode()
|
protectedvirtualinherited |
Get a leaf node.
Create a new node with the given id and name.
- Parameters
-
id The id of the node. name The name of the node.
- Returns
- A pointer toward a new node object.
Reimplemented in bpp::PGMA, and bpp::HierarchicalClustering.
Definition at line 111 of file AbstractAgglomerativeDistanceMethod.cpp.
◆ getName()
|
inlinevirtual |
- Returns
- The name of the distance method.
Implements bpp::DistanceMethod.
◆ getParentNode()
|
protectedvirtualinherited |
Get an inner node.
Create a new node with the given id, and set its sons.
- Parameters
-
id The id of the node. son1 The first son of the node. son2 The second son of the node.
- Returns
- A pointer toward a new node object.
Reimplemented in bpp::PGMA, and bpp::HierarchicalClustering.
Definition at line 116 of file AbstractAgglomerativeDistanceMethod.cpp.
References bpp::Node::addSon().
◆ getTree()
|
inlinevirtualinherited |
Get the computed tree, if there is one.
- Returns
- A copy of the computed tree if there is one, 0 otherwise.
Implements bpp::DistanceMethod.
Reimplemented in bpp::HierarchicalClustering, and bpp::PGMA.
Definition at line 127 of file AbstractAgglomerativeDistanceMethod.h.
References bpp::AbstractAgglomerativeDistanceMethod::tree_.
Referenced by bpp::TreeTools::MRP(), and bpp::TreeTools::MRPMultilabel().
◆ isVerbose()
|
inlinevirtualinherited |
- Returns
- True if verbose mode is enabled.
Implements bpp::DistanceMethod.
Definition at line 149 of file AbstractAgglomerativeDistanceMethod.h.
References bpp::AbstractAgglomerativeDistanceMethod::verbose_.
◆ outputPositiveLengths()
|
inlinevirtualinherited |
Definition at line 106 of file NeighborJoining.h.
References bpp::NeighborJoining::positiveLengths_.
Referenced by BioNJ().
◆ setDistanceMatrix()
|
inlinevirtual | ||||||||||||||
Set the distance matrix to use.
- Parameters
-
matrix The matrix to use.
- Exceptions
-
Exception In case an incorrect matrix is provided (eg smaller than 3).
Reimplemented from bpp::NeighborJoining.
Definition at line 101 of file BioNJ.h.
References bpp::NeighborJoining::setDistanceMatrix(), and variance_.
Referenced by BioNJ(), bpp::TreeTools::MRP(), and bpp::TreeTools::MRPMultilabel().
◆ setVerbose()
|
inlinevirtualinherited |
- Parameters
-
yn Enable/Disable verbose mode.
Implements bpp::DistanceMethod.
Definition at line 148 of file AbstractAgglomerativeDistanceMethod.h.
References bpp::AbstractAgglomerativeDistanceMethod::verbose_.
Member Data Documentation
◆ currentNodes_
|
protectedinherited |
Definition at line 69 of file AbstractAgglomerativeDistanceMethod.h.
Referenced by bpp::AbstractAgglomerativeDistanceMethod::operator=().
◆ lambda_
◆ matrix_
|
protectedinherited |
Definition at line 66 of file AbstractAgglomerativeDistanceMethod.h.
Referenced by bpp::AbstractAgglomerativeDistanceMethod::operator=().
◆ positiveLengths_
|
protectedinherited |
Definition at line 60 of file NeighborJoining.h.
Referenced by bpp::NeighborJoining::outputPositiveLengths().
◆ rootTree_
|
protectedinherited |
Definition at line 71 of file AbstractAgglomerativeDistanceMethod.h.
Referenced by bpp::AbstractAgglomerativeDistanceMethod::operator=().
◆ sumDist_
|
protectedinherited |
Definition at line 59 of file NeighborJoining.h.
Referenced by bpp::NeighborJoining::NeighborJoining(), and bpp::NeighborJoining::setDistanceMatrix().
◆ tree_
|
protectedinherited |
Definition at line 67 of file AbstractAgglomerativeDistanceMethod.h.
Referenced by bpp::AbstractAgglomerativeDistanceMethod::AbstractAgglomerativeDistanceMethod(), bpp::AbstractAgglomerativeDistanceMethod::getTree(), bpp::AbstractAgglomerativeDistanceMethod::operator=(), and bpp::AbstractAgglomerativeDistanceMethod::~AbstractAgglomerativeDistanceMethod().
◆ variance_
|
private |
Definition at line 59 of file BioNJ.h.
Referenced by setDistanceMatrix().
◆ verbose_
|
protectedinherited |
Definition at line 70 of file AbstractAgglomerativeDistanceMethod.h.
Referenced by bpp::AbstractAgglomerativeDistanceMethod::isVerbose(), bpp::AbstractAgglomerativeDistanceMethod::operator=(), and bpp::AbstractAgglomerativeDistanceMethod::setVerbose().
The documentation for this class was generated from the following files: